Lack of interpretability is a key obstacle in the adoption of ML models. Recent work on pre-miRNA detection has received good results. Interpreting the behavior of this model is yet to be developed. In this project, we aim to develop a deep generative model that can generate (simulate) miRNA molecules from latent factors with the goal of relating these factors to biological aspects.
MicroRNAs (miRNAs) are small non-coding RNA sequences that have been implicated in many physiological processes. Furthermore, miRNAs have been shown to be important biomarkers for diseases and their mimics are tested as drug candidates.
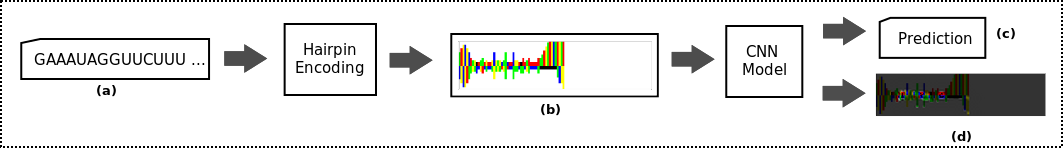
In previous work [1][2], we propose a method that uses domain knowledge to create an efficient image representation of miRNA molecules encoding sequence, structure, and implicitly some thermodynamic information. We then use this low-level feature representation of the molecules to develop a hierarchical deep representation using a convolutional neural network model, which directly detects precursor miRNAs. With this method, we achieve state-of-the-art performance on all previously used datasets.
The developed encoding and modeling process opens possibilities for interpretability of the models’ behavior, which may lead to novel biological interpretations of miRNA genesis and targeting. The goal of this project is to utilize these findings of the discriminative modeling for the development of a latent variable generative model such as conditional Variational Autoencoder that would allow us to generate miRNA molecules. Such a model would allow us to investigate the possibility to relate these latent factors to biologically meaningful structures or processes.
Supervisor: Vlado Menkovski
[1] J. Cordero, V. Menkovski, J. Allmeer. “Detection of pre-microRNA with Convolutional Neural Networks” (2019) bioRxiv 840579; doi: https://doi.org/10.1101/840579
[2] J. Cordero (2019) “Detection of pre-microRNAs with Convolutional Neural Networks” thesis
